Structure-guided protein engineering and immobilization of an amidohydrolase for efficient ochratoxin A detoxification.
Hu, Y., Xu, Y., He, B., Niu, D., Yang, X., Huang, J., Xie, Z., Shen, P., Li, X., Bai, M., Liu, Z., Li, H., Yang, Y., Huang, J.W., Chen, C.C., Guo, R.T., Dai, L.(2026) Int J Biol Macromol 358: 151671-151671
- PubMed: 41903634 Search on PubMed
- DOI: https://doi.org/10.1016/j.ijbiomac.2026.151671
- Primary Citation Related Structures:
9L2O, 9L2T, 9L6P - PubMed Abstract:
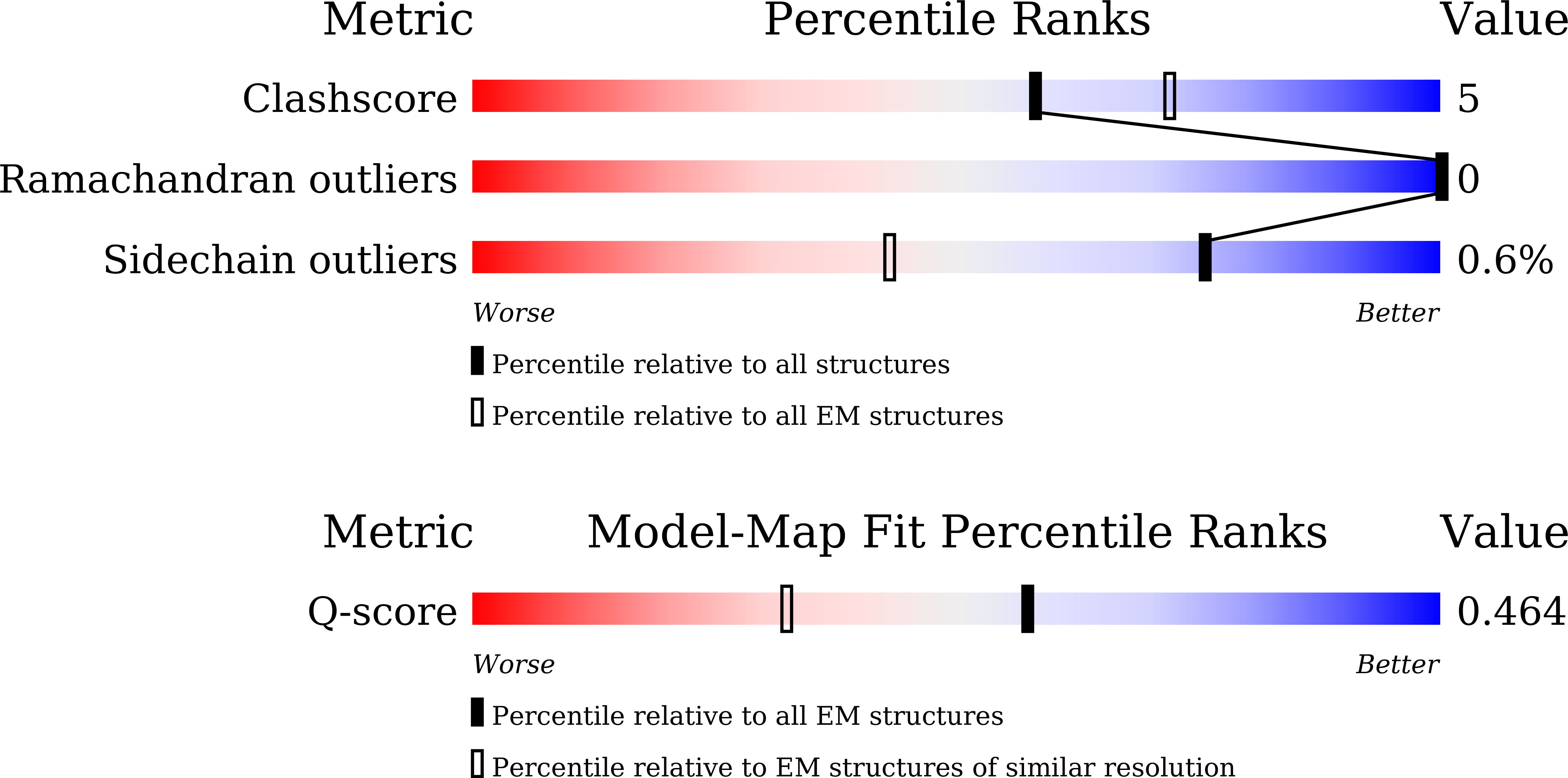
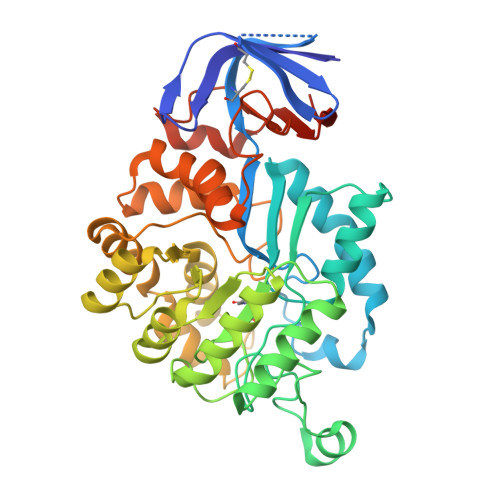
Ochratoxin A (OTA) is a pervasive and significant mycotoxin that poses serious health risks to humans and animals. The development of efficient biocatalysts for the enzymatic detoxification of OTA is of great importance. In this study, we enhanced the OTA-degrading activity of three amidohydrolases (ADHs) by up to ninefold. This improvement was achieved by reducing steric hindrance in the binding pocket and fine-tuning the hydrophilic interactions between the enzyme and substrate. The most efficient variant, PwADH/DM, was immobilized onto magnetic Fe 3 O 4 nanoparticles functionalized by the co-deposition of dopamine and polyethyleneimine. Under optimal conditions, this immobilization process achieved a high immobilization efficiency (85.4%) and activity recovery (76.9%). The immobilized enzyme exhibited enhanced pH stability and thermostability, along with good storage stability, reusability, and recyclability. More importantly, the immobilized enzyme completely degraded 100 ng/mL OTA in contaminated milk without affecting the milk's properties. These findings expand our understanding of the molecular mechanisms governing substrate binding and catalysis in OTA-degrading ADHs. Furthermore, they provide a blueprint for enzyme-based OTA decontamination during food processing.
- State Key Laboratory of Biocatalysis and Enzyme Engineering, Hubei Hongshan Laboratory, Hubei Key Laboratory of Industrial Biotechnology, School of Life Sciences, Hubei University, Wuhan, 430062, PR China.
Organizational Affiliation: